|
We have established methods for the discovery of drugs for treatment of American Trypanosomiasis (Chagas disease), using farnesyl pyrophosphate synthase from Trypanosoma cruzi as a target. To enable the discovery of drugs, it is essential to have access to relevant forms of the target protein and valid biochemical methods for studying the protein and effects of compounds that may be evolved into drugs. Particle concentration, type of ions in the medium can. This pilot screen confirms that the procedures developed herein enables SPR‐biosensor driven fragment‐based discovery of leads targeting tcFPPS, despite the lack of a reference compound. DLS can be used to characterize the size of various particles including proteins and small compounds. Instead, ranking could be performed from the slope of the linear relationship at low concentrations. Characterization of hits by SPR showed that all had low affinities and the relationships between steady‐state responses and concentrations were not sufficiently hyperbolic for determination of KD‐values. M > 1☌ and selectivity for tcFPPS in the presence of Mg2+. Dynamiclight(SLS) molecular weight scattering Electrophoreticlight Applicationexamples Molecularweight Sizing Polydispersity MalvernInstruments& (DLS)polydispersity scattering(ELS)zetapotential theZetasizerNano Thescatteringintensityisafunctionofthe olecularweightandconcentration. Hits were selected on the basis of response levels or ΔT A library of 90 fragments was efficiently screened by both assays at a single concentration in the presence and absence of the catalytic cofactor Mg2+. A thermal shift assay was used in parallel. An SPR‐biosensor assay for fragment library screening and hit confirmation was developed. 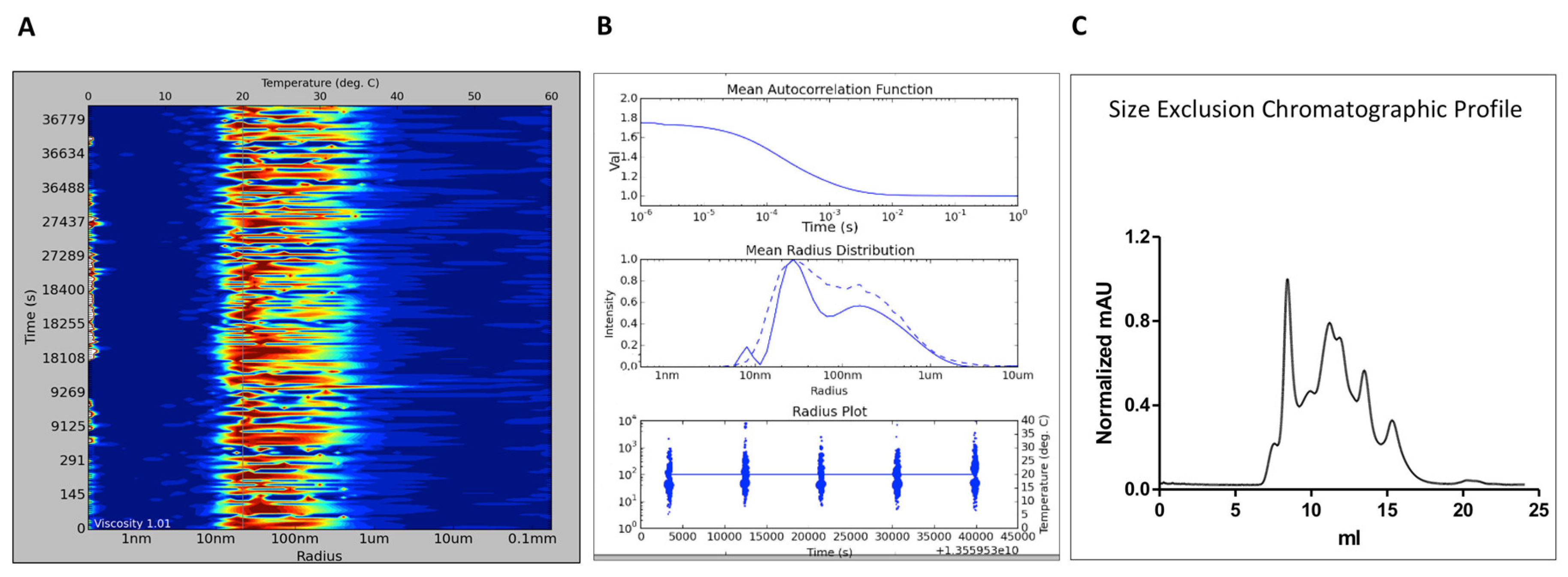
It also confirmed that sensor surfaces prepared with structurally intact protein was active. A new coupled enzymatic assay, using luciferase for detection of pyrophosphate, was developed and used to confirm that the purified enzyme was active after purification and storage. The improved methods enabled the production of pure, folded and dimeric protein, and identified procedures for storage and handling. Initial problems with protein stability and lack of useful reference compounds motivated optimization of experimental procedures and conditions. The method requires functional sensor surfaces with high sensitivity for extended times and appropriate controls. Here, in Part One of this series, we provide an overview of the key principles of DLS: theory, correlation statistics, deconvolution algorithms, and the intensity to mass transform.Procedures for producing and exploring Trypanosoma cruzi farnesyl pyrophosphate synthase (tcFPPS) for surface plasmon resonance (SPR) biosensor‐driven fragment‐based discovery have been established. This four-part white paper series examines common issues and questions surrounding the principles, measurements and analysis of DLS data and discusses how to minimize the time required for and increase the accuracy of acquiring and interpreting DLS data during the biotherapeutic development process. 
With that said, the challenge that remains with the application of DLS to protein therapeutic formulations is centered around data interpretation. The popularity of DLS within the biopharmaceutical industry is a consequence of the technique’s wide working size and extended sample concentration ranges, as well as its low volume requirements. Dynamic light scattering (DLS) is an analytical technique used to measure the particle size distribution of protein formulations across the oligomer and sub-micron size ranges of approximately 1 nm to 1 µm. Dynamic light scattering (DLS), also known as photon correlation spectroscopy (PCS), is a technique for determining the size distribution of vesicles.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
